|
We're hiring scientists & engineers
|
|
Welcome to the Allen Cell Explorer where you can access an unprecedented view into the organizational diversity of human stem cells through open data, tools, cell lines, plasmids, and models developed by the Allen Institute for Cell Science. All our scientific resources are openly available to support the cell biology community and accelerate discovery in human health and disease.
Highlights
Register for TEMTIA 11 |
|
We are excited to host the 11th EMT International Association (TEMTIA 11) meeting at the Allen Institute in Seattle, Washington, USA, on November 12-15, 2024.
|
The epithelial to mesenchymal transition (EMT) is a cell state change during which cells transition from stationary to migratory behavior. Associated with normal development but also a characteristic of many cancers, EMT includes complex changes across multiple cell characteristics and is consistently and universally difficult to define. At this year’s TEMTIA, we will leverage the diversity of the field, convening experts across contexts, perspectives, and scales of study to address this challenge. Scientific sessions will be organized with common themes such as: data integration, microscopy in EMT, modeling of state changes, and more.
To recognize the seminal contributions of Dr. Elizabeth Hay to the EMT field, TEMTIA will present the Betty Hay award of AU$1000.00 to a young female scientist. To apply for the fellowship or for more information, visit the Event Details page. Talk/poster abstract submission deadline: May 31, 2024 Please contact events@alleninstitute.org for any questions. |
New article published in Biophysical JournalWe just published a new Biophysical Perspective proposing an ambitious future for Cell Biology in the Biophysical Journal:
|
New article published in NatureDatabase of 200,000 cell images yields new mathematical framework to understand our cellular building blocks.
|
|
Featured Publications
|
Integrated intracellular organization and its variations in human iPS cells
Viana MP, Chen J, Knijnenburg TA, Vasan R, Yan C, Arakaki JE, Bailey M, Berry B, Borensztejn A, Brown EM, Carlson S, Cass JA, Chaudhuri B, Metzler KRC, Coston ME, Crabtree ZJ, Davidson S, DeLizo CM, Dhaka S, Dinh SQ, Do TP, Domingus J, Donovan-Maiye RM, Ferrante AJ, Foster TJ, Frick CL, Fujioka G, Fuqua MA, Gehring JL, Gerbin KA, Grancharova T, Gregor BW, Harrylock L, Haupt A, Hendershott MC, Hookway C, Horwitz AR, Hughes C, Isaac EJ, Johnson GR, Kim B, Leonard AN, Leung W, Lucas JJ, Ludmann SA, Lyons BM, Malik H, McGregor R, Medrash GE, Meharry SL, Mitcham K, Mueller IA, Murphy-Stevens TL, Nath A, Nelson AM, Oluoch SA, Paleologu L, Popiel TA, Riel-Mehan MM, Roberts B, Schaefbauer LM, Schwarzl M, Sherman J, Slaton S, Sluzewski MF, Smith JE, Sul Y, Swain-Bowden MJ, Tang WJ, Thirstrup DJ, Toloudis DT, Tucker AP, Valencia V, Wiegraebe W, Wijeratna T, Yang R, Zaunbrecher RJ, Labitigan RLD, Sanborn AL, Johnson GT, Gunawardane RN, Gaudreault N, Theriot JA, Rafelski SM. Nature, January 4, 2023. |
|
A comprehensive analysis of gene expression changes in a high replicate and open-source dataset of differentiating hiPSC-derived cardiomyocytes
Grancharova T, Gerbin KA, Rosenberg AB, Roco CM, Arakaki JE, DeLizo CM, Dinh SQ, Donovan-Maiye RM, Hirano M, Nelson AM, Tang J, Theriot JA, Yan C, Menon V, Palecek SP, Seelig G, Gunawardane RN. Scientific Reports, August 04, 2021. |
|
Cell states beyond transcriptomics: Integrating structural organization and gene expression in hiPSC-derived cardiomyocytes
Gerbin KA, Grancharova T, Donovan-Maiye RM, Hendershott MC, Anderson HG, Brown JM, Chen J, Dinh SQ, Gehring JL, Johnson GR, , Lee H, Nath A, Nelson AM, Sluzewski MF, Viana MP, Yan C, Zaunbrecher RJ, Cordes Metzler KR, Gaudreault N, Knijnenburg T, Rafelski SM, Theriot JA, Gunawardane RN. Cell Systems, June 16, 2021. |
We study the human cell holistically. To understand and predict cell behaviors, we develop and image high quality human induced pluripotent stem (hiPS) cell lines with genome editing to illuminate over 40 key cellular structures. This live cell imaging, together with image analysis, visualization tools and computational models enable novel interpretations of cell behavior by us and researchers around the globe.
Cells & plasmids for your lab
We have created a series of gene edited human iPSCs that we use as the foundation for our image data, and that we share with the larger scientific community.
|
Obtain cell lines & plasmids for your lab; quality control details
|
Explore & download genomics & transcriptomics data
|
Step-by-step protocols to culture & image gene-edited hiPSCs successfully
|
Instructional videos provided to ensure success in your lab
|
Featured digital tools
|
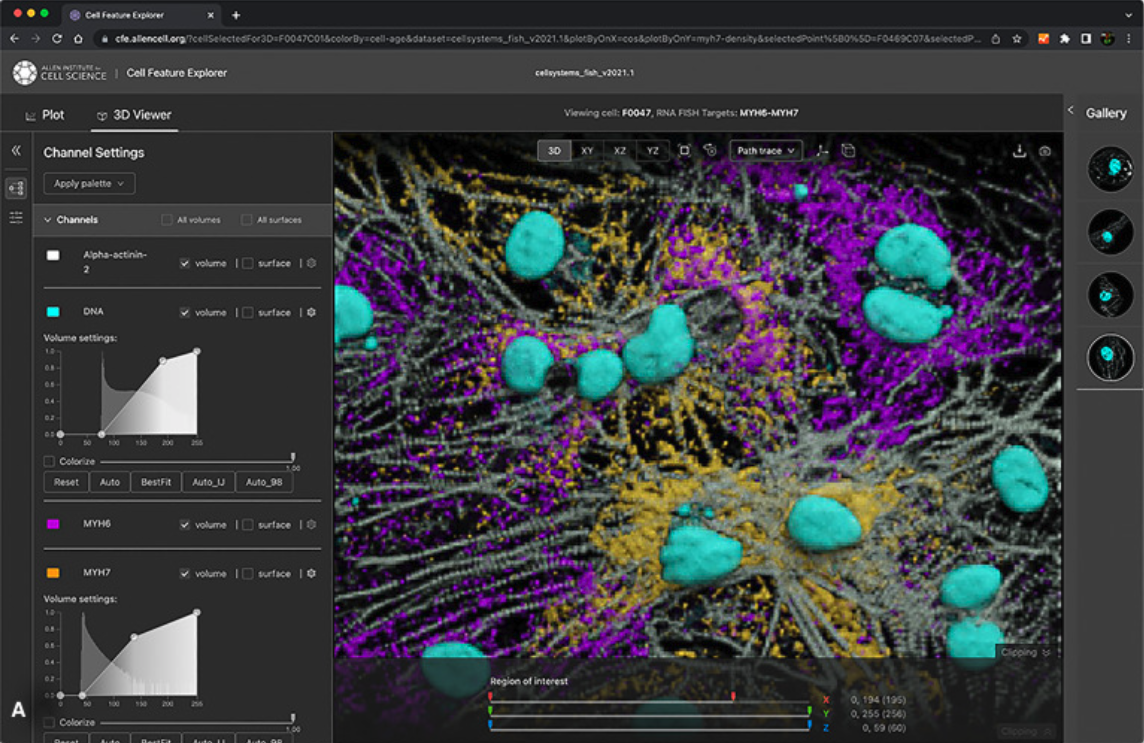
View multi-channel images using cinematographic rendering
|
Access & interact with our public 3D image datasets & feature measurements online
|
Perform 3D segmentations of intracellular structures
|
Access data & code developed at the institute
|